The Machine Learning for Ecology Research Group harnesses the power of machine learning, particularly deep learning, to advance research and discoveries in conservation ecology. With biodiversity declining at unprecedented rates, effective wildlife monitoring is more critical than ever to guide conservation efforts. Our primary goal is to improve wildlife monitoring and make ecological data analysis faster and more accessible. We believe that combining mathematics and cutting-edge technology offers powerful tools to deepen our understanding of biodiversity and ultimately support its protection.
Current Research Topics
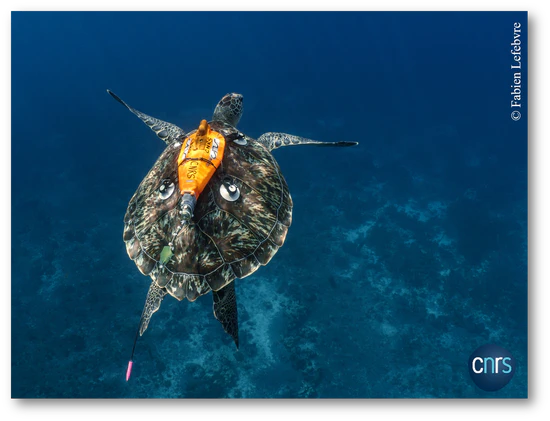
Biologgers are animal-borne devices that record high-frequency data (20–100 Hz) on both the animal and its environment. They can capture detailed information on posture and movement, providing valuable insights into animal behavior. When combined with labeled data, often obtained through animal-borne cameras, these recordings enable the training of deep learning algorithms to automatically identify behaviors.
The Machine Learning for Ecology research group focuses on developing such algorithms, applying them to wild populations, and enhancing their generalization through approaches such as transfer learning. The group also works on creating lightweight and efficient models that can be embedded directly on biologgers to enable real-time monitoring. This research is conducted in collaboration with the French National Centre for Scientific Research (CNRS) using data collected from sea turtles and with University of Cape town using data from chinstrap penguins
The Machine Learning for Ecology research group focuses on developing such algorithms, applying them to wild populations, and enhancing their generalization through approaches such as transfer learning. The group also works on creating lightweight and efficient models that can be embedded directly on biologgers to enable real-time monitoring. This research is conducted in collaboration with the French National Centre for Scientific Research (CNRS) using data collected from sea turtles and with University of Cape town using data from chinstrap penguins
Passive Acoustic Monitoring provides an effective approach to studying elusive wildlife by deploying acoustic recorders across multiple sites, often in remote locations, over extended periods. While this technique enables long-term, non-invasive monitoring, it also generates large volumes of data whose analysis can be both labor-intensive and time-consuming.
Deep learning has emerged as a powerful solution to this challenge, enabling the automated processing of acoustic data and the reliable detection of target species. In this context, our work focuses on developing deep learning tools to improve the efficiency and accuracy of species detection and classification from acoustic recordings.
Deep learning has emerged as a powerful solution to this challenge, enabling the automated processing of acoustic data and the reliable detection of target species. In this context, our work focuses on developing deep learning tools to improve the efficiency and accuracy of species detection and classification from acoustic recordings.
The African Penguin is an endangered species of penguin found only off the coast of Southern Africa. The aim of this project is to build a deep learning algorithm that can accurately predict penguin posture and depth from monocular video feed. This will allow us to classify penguin behaviours and better understand interactions between penguins. This will allow ecologists to gain better insights into penguin colonies and aid in conservation of this rapidly declining species.
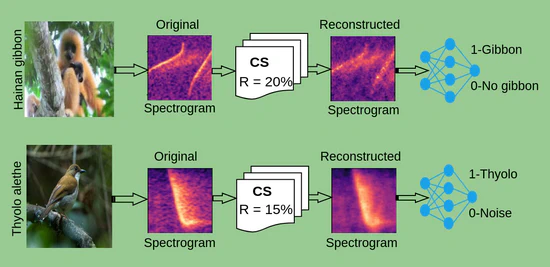
In this research, we investigate the potential of compressed sensing (CS) techniques for data compression as a step toward real-time bioacoustics monitoring. CS enables data to be compressed directly on recording devices, allowing efficient transmission over wireless networks or through low-bandwidth internet connections in remote areas. We compare CS with existing acoustic compression methods, with the goal of providing a framework to guide users in selecting the most suitable approach depending on the species under study.
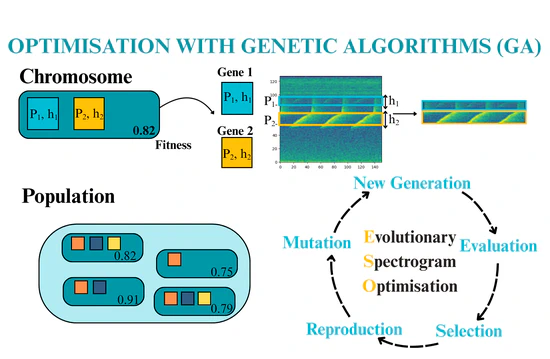
Passive Acoustic Monitoring (PAM) is a vital tool for wildlife monitoring, and deep learning, particularly convolutional neural networks (CNNs), has become the standard approach for developing bioacoustics classifiers. However, real-time classification remains challenging due to the high computational complexity of these models and the need for resource-intensive preprocessing steps, such as downsampling. One approach to reducing model size and computational complexity is decreasing input dimensions, specifically by reducing spectrogram size—the visual representation of sound. We introduce Evolutionary Spectrogram Optimisation (ESO), an evolutionary algorithm that automatically identifies and extracts narrow frequency bands from spectrograms that are informative enough to allow CNN-based species detection. ESO optimises the selection and size of these bands, while maximising classification accuracy and minimising the number of CNN parameters.

Latest Publications
- Jeantet L., Zondo K., Delvenne C., Martin J., Chevallier D., Dufourq E. (2024). Automatic identification of the endangered Hawksbill sea turtle behavior using deep learning and cross-species transfer learning. Journal of Experimental Biology. Doi: 10.1242/jeb.249232
- Hoffman B., Cusimano M., Baglione V., Canestrari D., Chevallier D., DeSantis D.L., Jeantet L. et al. (2024). A benchmark for computational analysis of animal behavior, using animal‑borne tags. Movement Ecology. Doi: 10.1186/s40462-024-00511-8
- Herbst C., Jeantet L., Dufourq E. (2024) Empirical Evaluation of Variational Autoencoders and Denoising Diffusion Models for Data Augmentation in Bioacoustics Classification. South African Computer Science and Information Systems Research Trends. Doi: 10.1007/978-3-031-64881-6_3
- Schoombie S., Jeantet L., Chimienti M., Sutton G.J., Pistorius P.A., Dufourq E., Lowther A.D., and Oosthuizen W.C. (2024). Identifying prey capture events of a free-ranging marine predator using bio-logger data and deep learning. Royal Society Open Science. Doi : 10.1098/rsos.240271